Off-the-shelf CAR-T cell therapies
CAR-T cell therapies have shown great promise in treating blood cancers and autoimmune diseases. Our CAR-T cell therapies are armored for potentially improved activity against diseases and are designed to reach a broad number of patients with off-the-shelf treatment options.
Scientific publications
Recent publications
 Encore: A CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (CB-010) for relapsed/refractory B cell non-Hodgkin lymphoma (r/r B-NHL): Updated phase 1 results from the ANTLER trial
Encore: A CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (CB-010) for relapsed/refractory B cell non-Hodgkin lymphoma (r/r B-NHL): Updated phase 1 results from the ANTLER trial
European Hematology Association (EHA) 2024 Hybrid Congress, June 13, 2024.
 Cas12a CRISPR Hybrid RNA-DNA (chRDNA)-Mediated In Vivo Genome-Editing Technology for Efficient and Functional Hepatic Gene Disruption
Cas12a CRISPR Hybrid RNA-DNA (chRDNA)-Mediated In Vivo Genome-Editing Technology for Efficient and Functional Hepatic Gene Disruption
American Society of Cell & Gene Therapy (ASGCT) Annual Meeting, May 7-11, 2024.
 CB-010 ANTLER Phase 1 trial: Results from dose escalation phase
CB-010 ANTLER Phase 1 trial: Results from dose escalation phase
iwCAR-T Meeting. April 20, 2024.
 Preclinical evaluation of CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy, that exhibits specific and potent cytotoxicity in acute myeloid leukemia (AML) xenograft models
Preclinical evaluation of CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy, that exhibits specific and potent cytotoxicity in acute myeloid leukemia (AML) xenograft models
American Association for Cancer Research (AACR) Annual Meeting. April 9, 2024.
 Improved genome editing with Cas12a and Cas9 chRDNA platform in T, NK, B, and iPS cells
Improved genome editing with Cas12a and Cas9 chRDNA platform in T, NK, B, and iPS cells
Transplantation & Cellular Therapy Meetings of ASTCT™ and CIBMTR®. February 21-24, 2024.
 High-specificity CRISPR-mediated genome engineering in anti-BCMA allogeneic CAR T cells suppresses allograft rejection in preclinical models
High-specificity CRISPR-mediated genome engineering in anti-BCMA allogeneic CAR T cells suppresses allograft rejection in preclinical models
Cancer Immunology Research. February 9, 2024.
 Deep and durable response in a patient with primary refractory DLBCL treated with CB-010, CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (Antler Trial)
Deep and durable response in a patient with primary refractory DLBCL treated with CB-010, CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (Antler Trial)
EBMT/EHA CAR T Cell Meeting. February 15-17, 2024.
 Durable complete response achieved in a relapsed/refractory diffuse large B cell lymphoma (DLBCL) patient treated with a CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout: Case report from the CB-010 ANTLER trial
Durable complete response achieved in a relapsed/refractory diffuse large B cell lymphoma (DLBCL) patient treated with a CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout: Case report from the CB-010 ANTLER trial
Lymphoma - Leukemia & Myeloma Congress. October 18-23, 2023.
 Conformational control of Cas endonucleases by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells
Conformational control of Cas endonucleases by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells
Annual Meeting of the American Society for Gene and Cell Therapy (ASGCT). May 16, 2022.
 A first-in-human Phase 1, multicenter, open-label study of CB-011, a next-generation CRISPR-genome edited allogeneic anti-BCMA immune-cloaked CAR-T cell therapy, in patients with relapsed/refractory multiple myeloma (CaMMouflage trial)
A first-in-human Phase 1, multicenter, open-label study of CB-011, a next-generation CRISPR-genome edited allogeneic anti-BCMA immune-cloaked CAR-T cell therapy, in patients with relapsed/refractory multiple myeloma (CaMMouflage trial)
American Society of Clinical Oncology (ASCO) Annual Meeting. June 5, 2023.
chRDNA and CRISPR
 Cas12a CRISPR Hybrid RNA-DNA (chRDNA)-Mediated In Vivo Genome-Editing Technology for Efficient and Functional Hepatic Gene Disruption
Cas12a CRISPR Hybrid RNA-DNA (chRDNA)-Mediated In Vivo Genome-Editing Technology for Efficient and Functional Hepatic Gene Disruption
American Society of Cell & Gene Therapy (ASGCT) Annual Meeting, May 7-11, 2024.
 Improved genome editing with Cas12a and Cas9 chRDNA platform in T, NK, B, and iPS cells
Improved genome editing with Cas12a and Cas9 chRDNA platform in T, NK, B, and iPS cells
Transplantation & Cellular Therapy Meetings of ASTCT™ and CIBMTR®. February 21-24, 2024.
 Conformational control of Cas endonucleases by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells
Conformational control of Cas endonucleases by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells
Annual Meeting of the American Society for Gene and Cell Therapy (ASGCT). May 16, 2022.
 Conformational control of Cas9 by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells
Conformational control of Cas9 by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells
Molecular Cell. September 2, 2021.
 Harnessing type I CRISPR–Cas systems for genome engineering in human cells
Harnessing type I CRISPR–Cas systems for genome engineering in human cells
Nature Biotechnology. November 18, 2019.
 Advances in industrial biotechnology using CRISPR-Cas systems
Advances in industrial biotechnology using CRISPR-Cas systems
Trends in Biotechnology. February 1, 2018.
 Mapping the genomic landscape of CRISPR-Cas9 cleavage
Mapping the genomic landscape of CRISPR-Cas9 cleavage
Nature Methods. May 1, 2017.
 DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks
DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks
Molecular Cell. August 4, 2016.
 Guide RNA functional modules direct Cas9 activity and orthogonality
Guide RNA functional modules direct Cas9 activity and orthogonality
Molecular Cell. October 16, 2014.
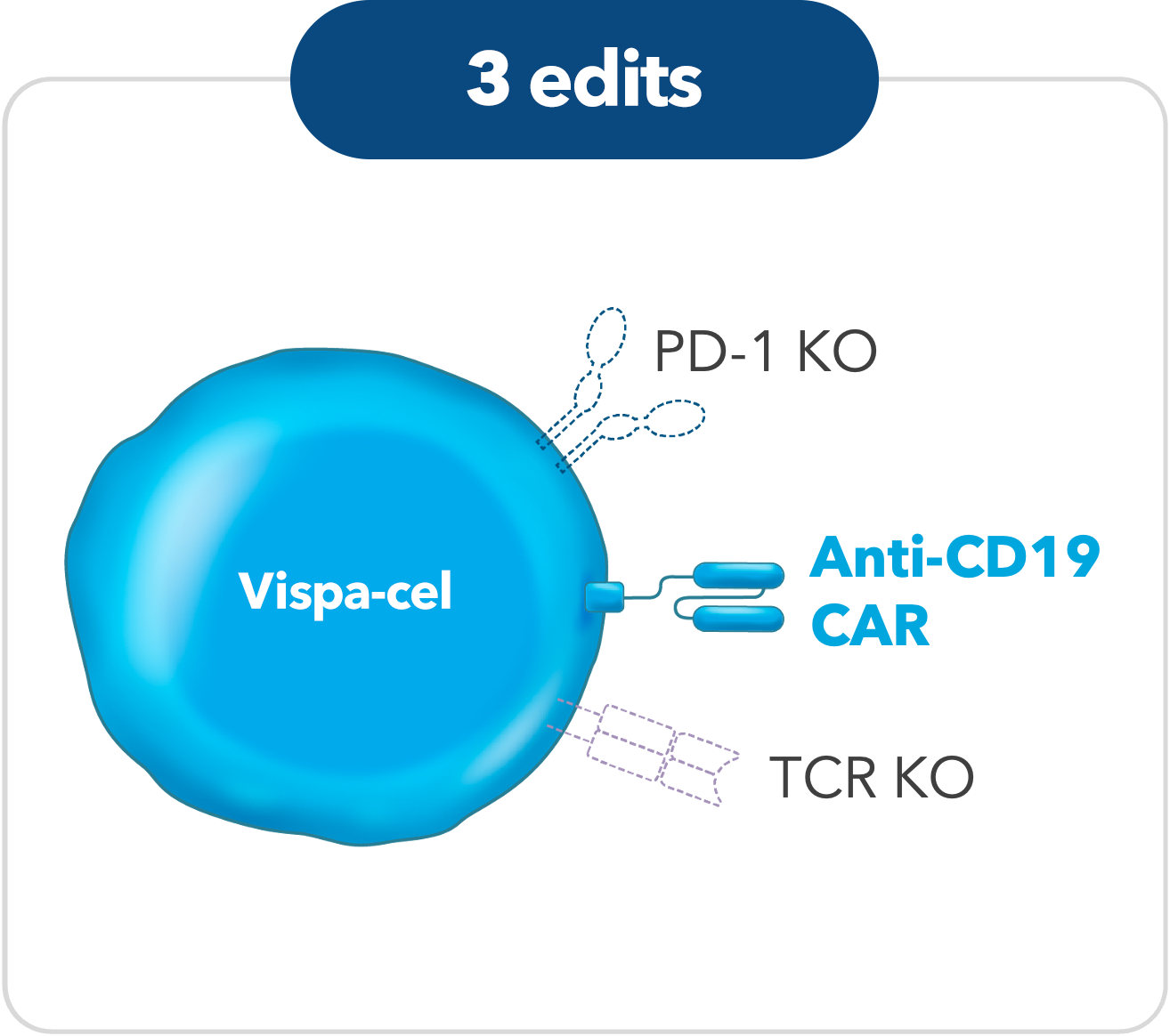
Vispa-cel
 Encore: A CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (CB-010) for relapsed/refractory B cell non-Hodgkin lymphoma (r/r B-NHL): Updated phase 1 results from the ANTLER trial
Encore: A CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (CB-010) for relapsed/refractory B cell non-Hodgkin lymphoma (r/r B-NHL): Updated phase 1 results from the ANTLER trial
European Hematology Association (EHA) 2024 Hybrid Congress, June 13, 2024.
 CB-010 ANTLER Phase 1 trial: Results from dose escalation phase
CB-010 ANTLER Phase 1 trial: Results from dose escalation phase
iwCAR-T Meeting. April 20, 2024.
 Deep and durable response in a patient with primary refractory DLBCL treated with CB-010, CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (Antler Trial)
Deep and durable response in a patient with primary refractory DLBCL treated with CB-010, CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy with a PD-1 knockout (Antler Trial)
EBMT/EHA CAR T Cell Meeting. February 15-17, 2024.
 Allogeneic chimeric antigen receptor-T cells with CRISPR-disrupted programmed death-1 checkpoint exhibit enhanced functional fitness
Allogeneic chimeric antigen receptor-T cells with CRISPR-disrupted programmed death-1 checkpoint exhibit enhanced functional fitness
Cytotherapy. Published online April 21, 2023.
 A First-in-Human Phase 1, Multicenter, Open-label Trial of CB-010, a Next-Generation CRISPR-Edited Allogeneic Anti-CD19 CAR-T Cell Therapy with a PD-1 Knockout, in Patients with Relapsed/Refractory B cell Non-Hodgkin Lymphoma (ANTLER Trial)
A First-in-Human Phase 1, Multicenter, Open-label Trial of CB-010, a Next-Generation CRISPR-Edited Allogeneic Anti-CD19 CAR-T Cell Therapy with a PD-1 Knockout, in Patients with Relapsed/Refractory B cell Non-Hodgkin Lymphoma (ANTLER Trial)
64th American Society of Hematology (ASH) Annual Meeting. December 10-13, 2022.
 CRISPR-edited Allogeneic Anti-CD19 CAR-T Cell Therapy with PD-1 Knockout Induces Prolonged Complete Response in Relapsed/Refractory Follicular Lymphoma Patient: Case Report from CB-010 ANTLER Trial
CRISPR-edited Allogeneic Anti-CD19 CAR-T Cell Therapy with PD-1 Knockout Induces Prolonged Complete Response in Relapsed/Refractory Follicular Lymphoma Patient: Case Report from CB-010 ANTLER Trial
Lymphoma, Leukemia, & Myeloma Congress. October 18-22, 2022.
 First-in-Human Trial of CB-010, a CRISPR-Edited Allogeneic Anti-CD19 CAR-T Cell Therapy with a PD-1 Knock Out, in Patients with Relapsed or Refractory B Cell Non-Hodgkin Lymphoma (ANTLER Study)
First-in-Human Trial of CB-010, a CRISPR-Edited Allogeneic Anti-CD19 CAR-T Cell Therapy with a PD-1 Knock Out, in Patients with Relapsed or Refractory B Cell Non-Hodgkin Lymphoma (ANTLER Study)
European Hematology Association (EHA) 2022 Hybrid Congress. June 10, 2022.
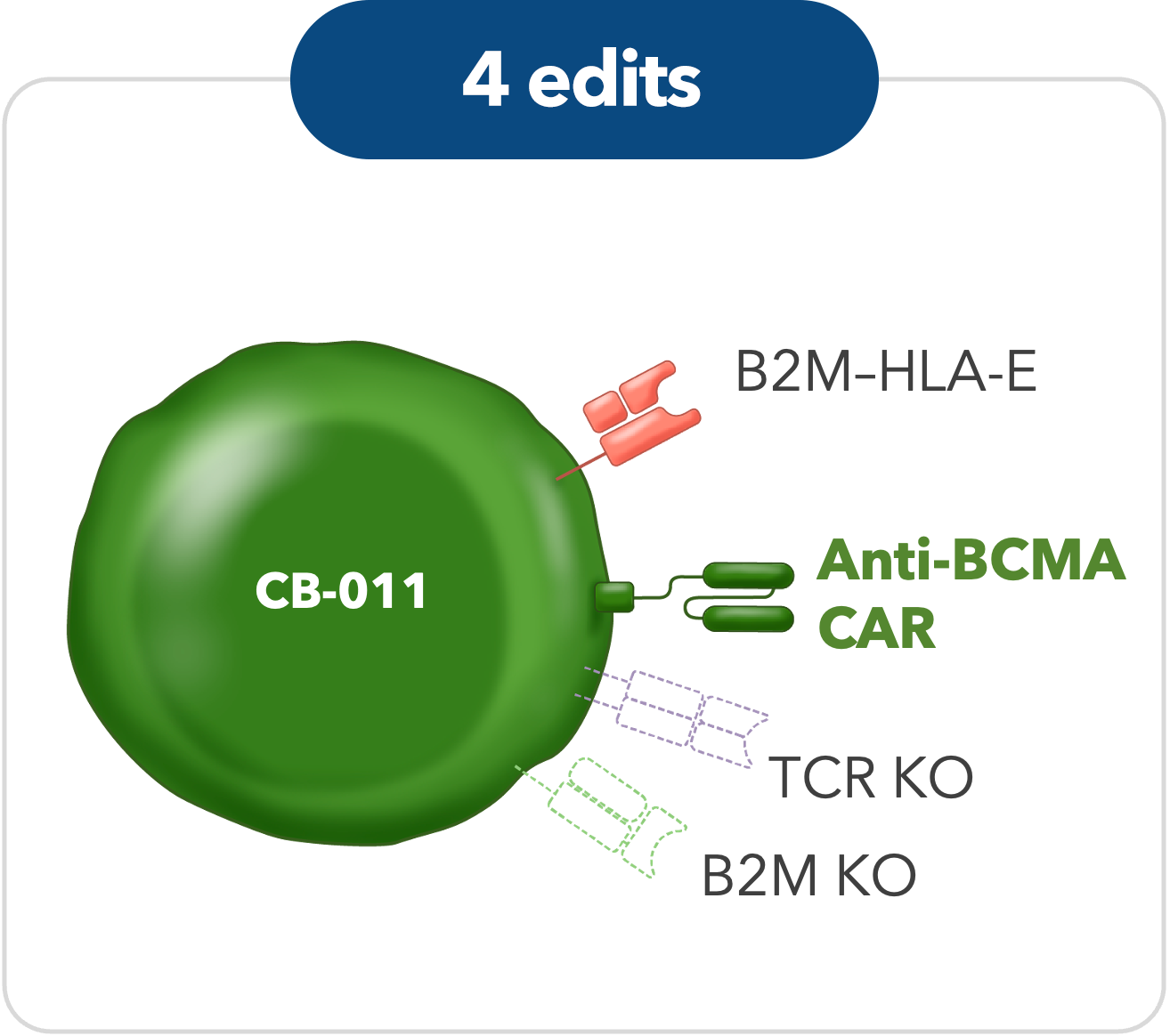
CB-011
 High-specificity CRISPR-mediated genome engineering in anti-BCMA allogeneic CAR T cells suppresses allograft rejection in preclinical models
High-specificity CRISPR-mediated genome engineering in anti-BCMA allogeneic CAR T cells suppresses allograft rejection in preclinical models
Cancer Immunology Research. February 9, 2024.
 A first-in-human Phase 1, multicenter, open-label study of CB-011, a next-generation CRISPR-genome edited allogeneic anti-BCMA immune-cloaked CAR-T cell therapy, in patients with relapsed/refractory multiple myeloma (CaMMouflage trial)
A first-in-human Phase 1, multicenter, open-label study of CB-011, a next-generation CRISPR-genome edited allogeneic anti-BCMA immune-cloaked CAR-T cell therapy, in patients with relapsed/refractory multiple myeloma (CaMMouflage trial)
American Society of Clinical Oncology (ASCO) Annual Meeting. June 5, 2023.
 CB-011, a BCMA-specific allogeneic CAR-T cell therapy, engineered with next-generation CRISPR technology to knock out B2M and express myelomar/r multiple transgene to blunt immune cell-mediated rejection, for B2M–HLA-E fusion
CB-011, a BCMA-specific allogeneic CAR-T cell therapy, engineered with next-generation CRISPR technology to knock out B2M and express myelomar/r multiple transgene to blunt immune cell-mediated rejection, for B2M–HLA-E fusion
TANDEM MEETINGS: Transplantation & Cellular Therapy Meetings of ASTCT™ and CIBMTR®. February 15-19, 2023.
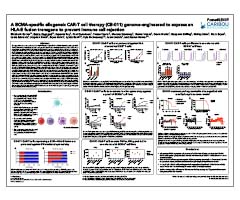 A BCMA-specific allogeneic CAR-T cell therapy (CB-011) genome-engineered to express an HLA-E fusion transgene to prevent immune cell rejection
A BCMA-specific allogeneic CAR-T cell therapy (CB-011) genome-engineered to express an HLA-E fusion transgene to prevent immune cell rejection
American Association for Cancer Research (AACR) Annual Meeting. April 10, 2022.
Other
 Preclinical evaluation of CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy, that exhibits specific and potent cytotoxicity in acute myeloid leukemia (AML) xenograft models
Preclinical evaluation of CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy, that exhibits specific and potent cytotoxicity in acute myeloid leukemia (AML) xenograft models
American Association for Cancer Research (AACR) Annual Meeting. April 9, 2024.
 CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy engineered with next-generation CRISPR technology to resist both the immunosuppressive tumor microenvironment and immune cell-mediated rejection, for patients with relapsed or refractory acute myeloid leukemia
CB-012, an allogeneic anti-CLL-1 CAR-T cell therapy engineered with next-generation CRISPR technology to resist both the immunosuppressive tumor microenvironment and immune cell-mediated rejection, for patients with relapsed or refractory acute myeloid leukemia
American Association for Cancer Research (AACR) Annual Meeting. April 17, 2023.
 CB-020, an Induced Pluripotent Stem Cell (iPSC)-Derived Allogeneic CAR-NK Cell Therapy, Engineered for Enhanced Activity Against Solid Tumors
CB-020, an Induced Pluripotent Stem Cell (iPSC)-Derived Allogeneic CAR-NK Cell Therapy, Engineered for Enhanced Activity Against Solid Tumors
12th AACR-JCA Joint Conference: Breakthroughs in Cancer Research – Translating Knowledge into Practice. December 10-14, 2022.








